Creating plots and graphs using the ggplot2 package.
Author
Thanks to Roy Francis, NBIS, RaukR2023 for sharing it
Note
These are a series of exercises to help you get started and familiarize yourself with ggplot2 syntax, plot building logic and fine modification of plots. Practice using the Basics section and then move on to slightly more complex plots: a scatterplot and a heatmap.
1. Basics
First step is to make sure that the necessary packages are installed and loaded.
Code
library(dplyr)
Attaching package: 'dplyr'
The following objects are masked from 'package:stats':
filter, lag
The following objects are masked from 'package:base':
intersect, setdiff, setequal, union
Code
library(tidyr)library(stringr)library(ggplot2)
Warning: package 'ggplot2' was built under R version 4.4.3
Code
library(ggrepel)library(patchwork)
Warning: package 'patchwork' was built under R version 4.4.3
We use the iris data to get started. This dataset has four continuous variables and one categorical variable. It is important to remember about the data type when plotting graphs.
Code
data("iris")head(iris)
Sepal.Length
Sepal.Width
Petal.Length
Petal.Width
Species
5.1
3.5
1.4
0.2
setosa
4.9
3.0
1.4
0.2
setosa
4.7
3.2
1.3
0.2
setosa
4.6
3.1
1.5
0.2
setosa
5.0
3.6
1.4
0.2
setosa
5.4
3.9
1.7
0.4
setosa
Building a plot
ggplot2 plots are initialized by specifying the dataset. This can be saved to a variable or it draws a blank plot.
Code
ggplot(data=iris)
Now we can specify what we want on the x and y axes using aesthetic mapping. And we specify the geometric using geoms. Note that the variable names do not have double quotes "" like in base plots.
Further geoms can be added. For example let’s add a regression line. When multiple geoms with the same aesthetics are used, they can be specified as a common mapping. Note that the order in which geoms are plotted depends on the order in which the geoms are supplied in the code. In the code below, the points are plotted first and then the regression line.
We can use the categorical column Species to color the points. The color aesthetic is used by geom_point and geom_smooth. Three different regression lines are now drawn. Notice that a legend is automatically created.
We can map another variable as size of the points. This is done by specifying size inside the aesthetic mapping. Now the size of the points denote Sepal.Width. A new legend group is created to show this new aesthetic.
Now let’s rename the axis labels, change the legend title and add a title, a subtitle and a caption. We change the legend title using scale_color_continuous(). All other labels are changed using labs().
Code
ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")
`geom_smooth()` using formula = 'y ~ x'
Axes modification
Let’s say we are not happy with the x-axis breaks 2,4,6 etc. We would like to have 1,2,3… We change this using scale_x_continuous().
Code
ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")
`geom_smooth()` using formula = 'y ~ x'
Faceting
We can create subplots using the faceting functionality. Let’s create three subplots for the three levels of Species.
Code
ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")+facet_wrap(~Species)
`geom_smooth()` using formula = 'y ~ x'
Themes
The look of the plot can be changed using themes. Let’s can the default theme_grey() to theme_bw().
Code
ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")+facet_wrap(~Species)+theme_bw()
`geom_smooth()` using formula = 'y ~ x'
All non-data related aspects of the plot can be modified through themes. Let’s modify the colors of the title labels and turn off the gridlines. The various parameters for theme can be found using ?theme.
Code
ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")+facet_wrap(~Species)+theme_bw()+theme(axis.title=element_text(color="Blue",face="bold"),plot.title=element_text(color="Green",face="bold"),plot.subtitle=element_text(color="Pink"),panel.grid=element_blank() )
`geom_smooth()` using formula = 'y ~ x'
Themes can be saved and reused.
Code
newtheme <-theme(axis.title=element_text(color="Blue",face="bold"),plot.title=element_text(color="Green",face="bold"),plot.subtitle=element_text(color="Pink"),panel.grid=element_blank())ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")+facet_wrap(~Species)+theme_bw()+ newtheme
`geom_smooth()` using formula = 'y ~ x'
Controlling legends
Here we see two legends based on the two aesthetic mappings.
Warning: ggrepel: 134 unlabeled data points (too many overlaps). Consider
increasing max.overlaps
Annotations
Custom annotations of any geom can be added arbitrarily anywhere on the plot.
Code
ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species))+annotate("text",x=2.5,y=2.1,label="There is a random line here")+annotate("segment",x=2,xend=4,y=1.5,yend=2)
An example of using error bars with points. The mean and standard deviation is computed. This is used to create upper and lower bounds for the error bars.
The aim of this challenge is to recreate the plot below originally published in The Economist. The graph is a scatterplot showing the relationship between Corruption Index and Human Development Index for various countries.
Make sure that the fields are of the correct type. The x-axis field ‘CPI’ and the y-axis field ‘HDI’ must be of numeric type. The categorical field ‘Region’ must be of Factor type.
Code
str(ec)
'data.frame': 173 obs. of 6 variables:
$ X : int 1 2 3 4 5 6 7 8 9 10 ...
$ Country : chr "Afghanistan" "Albania" "Algeria" "Angola" ...
$ HDI.Rank: int 172 70 96 148 45 86 2 19 91 53 ...
$ HDI : num 0.398 0.739 0.698 0.486 0.797 0.716 0.929 0.885 0.7 0.771 ...
$ CPI : num 1.5 3.1 2.9 2 3 2.6 8.8 7.8 2.4 7.3 ...
$ Region : chr "Asia Pacific" "East EU Cemt Asia" "MENA" "SSA" ...
We need to first modify the region column. The current levels in the ‘Region’ field are:
Code
levels(ec$Region)
NULL
But, the categories on the plot are different and need to be changed as follows:
From To
EU W. Europe OECD
Americas Americas
Asia Pacific Asia & Oceania
East EU Cemt Asia Central & Eastern Europe
MENA Middle East & North Africa
SSA Sub-Saharan Africa
Since the ‘To’ strings are a bit too long to be in one line on the legend, use \n to break a line into two lines.
Tip
\n is the newline character in R.
From To
EU W. Europe OECD
Americas Americas
Asia Pacific Asia &\nOceania
East EU Cemt Asia Central &\nEastern Europe
MENA Middle East &\nNorth Africa
SSA Sub-Saharan\nAfrica
The strings can be renamed using string replacement or substitution. But a easier way to do it is to use factor(). The arguments levels and labels in function factor() can be used to rename factors.
Code
ec$Region <-factor(ec$Region,levels =c("EU W. Europe","Americas","Asia Pacific","East EU Cemt Asia","MENA","SSA"),labels =c("OECD","Americas","Asia &\nOceania","Central &\nEastern Europe","Middle East &\nNorth Africa","Sub-Saharan\nAfrica"))
Provide data.frame ‘ec’ as the data and map field ‘CPI’ to the x-axis and ‘HDI’ to the y-axis. Use geom_point() to draw point geometry. To select shapes, see here. Circular shape can be drawn using 1, 16, 19, 20 and 21. Using shape ‘21’ allows us to control stroke color, fill color and stroke thickness for the points. Check out ?geom_point and look under ‘Aesthetics’ for the various possible aesthetic options. Set shape to 21, size to 3, stroke to 0.8 and fill to white.
Now, we add the trend line using geom_smooth. Check out ?geom_smooth and look under ‘Arguments’ for argument options and ‘Aesthetics’ for the aesthetic options.
Use method ‘lm’ and use a custom formula of y~poly(x,2) to approximate the curve seen on the plot. Turn off confidence interval shading. Set line thickness to 0.6 and line color to red.
Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
ℹ Please use `linewidth` instead.
Notice that the line in drawn over the points due to the plotting order. We want the points to be over the line. So reorder the geoms. Since we provided no aesthetic mappings to geom_smooth, there is no legend entry for the trendline. We can fake a legend entry by providing an aesthetic, for example; aes(fill="red"). We do not use the color aesthetic because it is already in use and would give us reduced control later on to modify this legend entry.
Code
p <-ggplot(ec,aes(x=CPI,y=HDI,color=Region))+geom_smooth(aes(fill="red"),method="lm",formula=y~poly(x,2),se=F,color="red",size=0.6)+geom_point(shape=21,size=3,stroke=0.8,fill="white")p
Text Labels
Now we add the text labels. Only a subset of countries are plotted. The list of countries to label is shown below.
Custom font can be used for the labels by providing the font name to argument family like so geom_text(family="fontname"). If you do not want to bother with fonts, just avoid the family argument in geom_text and skip this part.
Using custom fonts can be tricky business. To use a font name, it must be installed on your system and it should be imported into the R environment. This can be done using the extrafont package. Try importing one of the fonts available on your system. Not all fonts work. extrafont prefers .ttf fonts. If a font doesn’t work, try another.
Code
library(extrafont)font_import(pattern="Gidole",prompt=FALSE)# load fonts for pdfloadfonts()# list available fonts in Rfonts()
The actual font used on the Economist graph is something close to ITC Officina Sans. Since this is not a free font, I am using a free font called Gidole.
Warning in grid.Call.graphics(C_text, as.graphicsAnnot(x$label), x$x, x$y, :
font family not found in Windows font database
Label Overlap
To avoid overlapping of labels, we can use a ggplot2 extension package ggrepel. We can use function geom_text_repel() from the ggrepel package. geom_text_repel() has the same arguments/aesthetics as geom_text and a few more. Skip the family=Gidole part if you do not want to change the font.
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Axes
Next step is to adjust the axes breaks, axes labels, point colors and relabeling the trendline legend text.
Change axes labels to ‘Corruption Perceptions Index, 2011 (10=least corrupt)’ on the x-axis and ‘Human Development Index, 2011 (1=best)’ on the y-axis. Set breaks on the x-axis from 1 to 10 by 1 increment and y-axis from 0.2 to 1.0 by 0.1 increments.
Code
p <- p+scale_x_continuous(name="Corruption Perceptions Index, 2011 (10=least corrupt)",breaks=1:10,limits=c(1,10))+scale_y_continuous(name="Human Development Index, 2011 (1=best)",breaks=seq(from=0,to=1,by=0.1),limits=c(0.2,1))p
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Scale Colors
Now we want to change the color palette for the points and modify the legend text for the trendline.
Use scale_color_manual() to provide custom colors. These are the colors to use for the points: "#23576E","#099FDB","#29B00E", "#208F84","#F55840","#924F3E".
Use scale_fill_manual to change the trendline label since it’s a fill scale. The legend entry for the trendline should read ‘R^2=52%’.
Code
p <- p+scale_color_manual(values=c("#23576E","#099FDB","#29B00E", "#208F84","#F55840","#924F3E"))+scale_fill_manual(name="trend",values="red",labels=expression(paste(R^2,"=52%")))p
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Warning in grid.Call(C_textBounds, as.graphicsAnnot(x$label), x$x, x$y, : font
family not found in Windows font database
Title
Title and caption can be added with labs.
Set the title to ‘Corruption and human development’.
Set the caption to ‘Sources: Transparency International; UN Human Development Report’.
Code
p <- p+labs(title="Corruption and human development",caption="Sources: Transparency International; UN Human Development Report")p
Theme
We want to move the legend to the top and as a single row. This can be done using theme() option legend.position. See ?theme. guides() is used to set the number of rows to 1. We also set a custom font for all text elements using base_family="Gidole". This can be skipped if a font change is not required.
Code
p <- p+guides(color=guide_legend(nrow=1))+theme_bw(base_family="Gidole")+theme(legend.position="top")p
And now our plot is ready and we can compare with the original. Close? Close enough?
The full script for this challenge is summarized here:
Code
# read dataec <-read.csv("assets/data_economist.csv",header=T)# refactorec$Region <-factor(ec$Region,levels =c("EU W. Europe","Americas","Asia Pacific","East EU Cemt Asia","MENA","SSA"),labels =c("OECD","Americas","Asia &\nOceania","Central &\nEastern Europe","Middle East &\nNorth Africa","Sub-Saharan\nAfrica"))# labelslabels <-c("Congo","Afghanistan","Sudan","Myanmar","Iraq","Venezuela","Russia","Argentina","Brazil","Italy","South Africa","Cape Verde","Bhutan","Botswana","Britian","New Zealand","Greece","China","India","Rwanda","Spain","France","United States","Japan","Norway","Singapore","Barbados","Germany")# plottingp1 <-ggplot(ec,aes(x=CPI,y=HDI,color=Region))+geom_smooth(aes(fill="red"),method="lm",formula=y~poly(x,2),se=F,color="red",size=0.6)+geom_point(shape=21,size=3,stroke=0.8,fill="white")+geom_text_repel(data=subset(ec,Country %in% labels),aes(label=Country),color="black",box.padding=unit(1,'lines'),segment.size=0.25,size=3,family="Gidole")+scale_x_continuous(name="Corruption Perceptions Index, 2011 (10=least corrupt)",breaks=1:10,limits=c(1,10))+scale_y_continuous(name="Human Development Index, 2011 (1=best)",breaks=seq(from=0,to=1,by=0.1),limits=c(0.2,1))+scale_color_manual(values=c("#23576E","#099FDB","#29B00E", "#208F84","#F55840","#924F3E"))+scale_fill_manual(name="trend",values="red",labels=expression(paste(R^2,"=52%")))+labs(title="Corruption and human development",caption="Sources: Transparency International; UN Human Development Report")+guides(color=guide_legend(nrow=1))+theme_bw(base_family="Gidole")+theme(legend.position="top",panel.grid.minor=element_blank(),panel.grid.major.x=element_blank(),panel.background=element_blank(),panel.border=element_blank(),legend.title=element_blank(),axis.title=element_text(face="italic"),axis.ticks.y=element_blank(),axis.ticks.x=element_line(color="grey60"),plot.title=element_text(face="bold"),plot.caption=element_text(hjust=0,size=8))p1
3. WSJ Heatmap
The aim of this challenge is to recreate the plot below originally published in The Wall Street Journal. The plot is a heatmap showing the normalized number of cases of measles across 51 US states from 1928 to 2003. X-axis shows years and y-axis shows the names of states. The color of the tiles denote the number of measles cases per 100,000 people. Introduction of the measles vaccine is shown as the black line in 1963.
Start by reading in the data. This .csv file has two lines of comments so we need to skip 2 lines while reading in the data. We also add stringsAsFactors=F to avoid the automatic conversion of character fields to factor type.
Code
me <-read.csv("assets/data_wsj.csv",header=T,stringsAsFactors=F,skip=2)head(me)
YEAR state value
1 1928 ALABAMA 3.67
2 1929 ALABAMA 3.20
...
5501 1957 ALASKA 2.16
5502 1958 ALASKA 2.05
...
The solution is to sum up all the cases for a state for all weeks within a year into one value for that year. This can be done using the summarise() function from package dplyr.
A custom function is used to sum over weeks. If all values are NA, then result is NA. If some values are NA, the NAs are removed and the remaining numbers are summed.
The dots in state names are replaced by spaces and the words are converted to title case (First letter capital and rest lowercase).
We also convert the column names to lowercase for consistency.
We can build up a basic ggplot and heatmap tiles can be plotted using the geom geom_tile. ‘year’ is mapped to the x-axis, ‘state’ to the y-axis and fill color for the tiles is the ‘total’ value.
Add borders around the tiles. We use reorder(state,desc(state)) to reverse the order of states so that it reads A-Z from top to bottom.
Code
p <-ggplot(me3,aes(x=year,y=reorder(state,desc(state)),fill=total))+geom_tile(color="white",size=0.25)p
Scales
The extra space on left and right (gray) of the plot is removed using argument expand in scales. X-axis breaks are redefined at 10 year intervals from 1930 to 2010. Custom colors are used for the tiles: "#e7f0fa","#c9e2f6","#95cbee","#0099dc","#4ab04a", "#ffd73e","#eec73a","#e29421","#f05336","#ce472e". Since the color scale is a fill color on a continuous value and we want to supply n new colors, we use scale_fill_gradientn. Tiles with missing value is set to the color "grey90".
We can remove the x and y axes titles and add a plot title.
Code
p <- p+labs(x="",y="",fill="",title="Measles")p
Fixed Coords
We can use coord_fixed() to fix the coordinates for equal values in x and y direction. This should render perfectly square tiles.
Code
p <- p+coord_fixed()p
Annotation
Add the annotation line and text to denote the introduction of the vaccine. The line is at the position 1963. Custom font ‘Gidole’ is used here. This can be skipped.
Code
p <- p+geom_segment(x=1963,xend=1963,y=0,yend=51.5,size=.6,alpha=0.7) +annotate("text",label="Vaccine introduced",x=1963,y=53, vjust=1,hjust=0,size=I(3),family="Gidole")p
Warning in grid.Call.graphics(C_text, as.graphicsAnnot(x$label), x$x, x$y, :
font family not found in Windows font database
Theme
Here we change the following aspects of the plot using theme:
Change theme to theme_minimal to remove unnecessary plot elements.
Optional custom font. See ‘Custom font’ section under ‘Economist Scatterplot’.
Position the legend to bottom center.
Set legend font to color grey20.
Adjust size and justification of x and y axes text
---title: "ggplot2 Part 2"author: "Thanks to Roy Francis, NBIS, RaukR2023 for sharing it"description: "Creating plots and graphs using the ggplot2 package."image: "assets/featured.jpg"format: html: title-block-banner: true smooth-scroll: true toc: true toc-depth: 4 toc-location: right number-types: true number-depth: 4 code-fold: true code-tools: true code-copy: true code-overflow: wrap df-print: kable standalone: false fig-align: left theme: pulse highlight: kate---::: {.callout-note}These are a series of exercises to help you get started and familiarize yourself with ggplot2 syntax, plot building logic and fine modification of plots. Practice using the *Basics* section and then move on to slightly more complex plots: a scatterplot and a heatmap.:::<br>:::: {.columns}::: {.column width="50%"}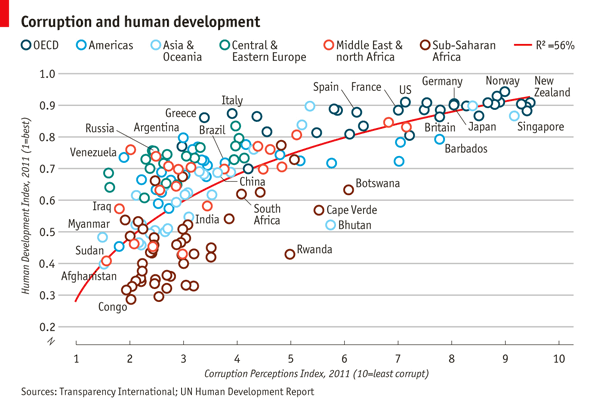:::::: {.column width="50%"}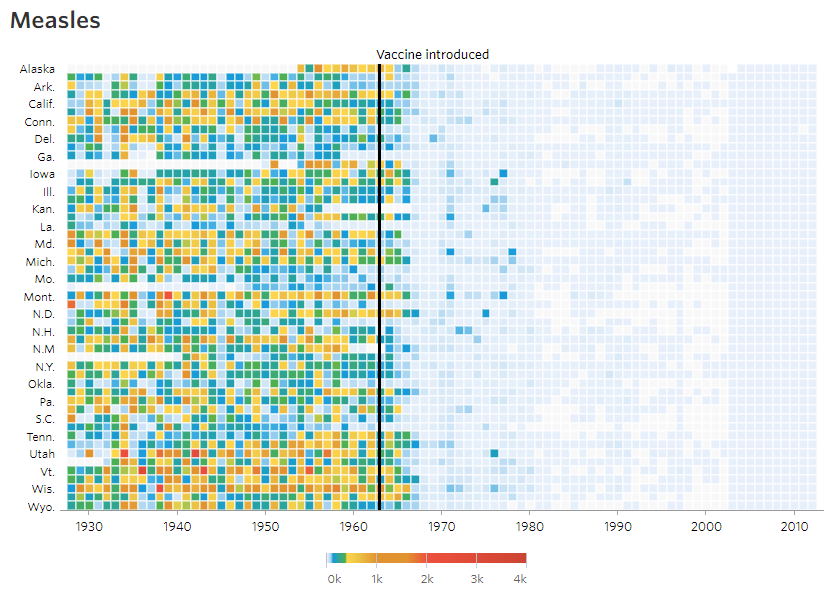:::::::<br>## 1. BasicsFirst step is to make sure that the necessary packages are installed and loaded.```{r}library(dplyr)library(tidyr)library(stringr)library(ggplot2)library(ggrepel)library(patchwork)```We use the `iris` data to get started. This dataset has four continuous variables and one categorical variable. It is important to remember about the data type when plotting graphs.```{r}data("iris")head(iris)```### Building a plotggplot2 plots are initialized by specifying the dataset. This can be saved to a variable or it draws a blank plot.```{r}#| fig-height: 4#| fig-width: 4ggplot(data=iris)```Now we can specify what we want on the x and y axes using aesthetic mapping. And we specify the geometric using geoms. Note that the variable names do not have double quotes `""` like in base plots.```{r}#| fig-height: 4#| fig-width: 4ggplot(data=iris)+geom_point(mapping=aes(x=Petal.Length,y=Petal.Width))```### Multiple geomsFurther geoms can be added. For example let's add a regression line. When multiple geoms with the same aesthetics are used, they can be specified as a common mapping. Note that the order in which geoms are plotted depends on the order in which the geoms are supplied in the code. In the code below, the points are plotted first and then the regression line.```{r}#| fig-height: 4#| fig-width: 4ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point()+geom_smooth(method="lm")```### Using colorsWe can use the categorical column `Species` to color the points. The color aesthetic is used by `geom_point` and `geom_smooth`. Three different regression lines are now drawn. Notice that a legend is automatically created.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width,color=Species))+geom_point()+geom_smooth(method="lm")```If we wanted to keep a common regression line while keeping the colors for the points, we could specify color aesthetic only for `geom_point`.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species))+geom_smooth(method="lm")```### Aesthetic parameterWe can change the size of all points by a fixed amount by specifying size outside the aesthetic parameter.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species),size=3)+geom_smooth(method="lm")```### Aesthetic mappingWe can map another variable as size of the points. This is done by specifying size inside the aesthetic mapping. Now the size of the points denote `Sepal.Width`. A new legend group is created to show this new aesthetic.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species,size=Sepal.Width))+geom_smooth(method="lm")```### Discrete colorsWe can change the default colors by specifying new values inside a scale.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species,size=Sepal.Width))+geom_smooth(method="lm")+scale_color_manual(values=c("red","blue","green"))```### Continuous colorsWe can also map the colors to a continuous variable. This creates a color bar legend item.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")```### TitlesNow let's rename the axis labels, change the legend title and add a title, a subtitle and a caption. We change the legend title using `scale_color_continuous()`. All other labels are changed using `labs()`.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")```### Axes modificationLet's say we are not happy with the x-axis breaks 2,4,6 etc. We would like to have 1,2,3... We change this using `scale_x_continuous()`.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")```### FacetingWe can create subplots using the faceting functionality. Let's create three subplots for the three levels of Species.```{r}#| fig-height: 4#| fig-width: 6ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")+facet_wrap(~Species)```### ThemesThe look of the plot can be changed using themes. Let's can the default `theme_grey()` to `theme_bw()`.```{r}#| fig-height: 4#| fig-width: 6ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")+facet_wrap(~Species)+theme_bw()```All non-data related aspects of the plot can be modified through themes. Let's modify the colors of the title labels and turn off the gridlines. The various parameters for theme can be found using `?theme`.```{r}#| fig-height: 4#| fig-width: 6ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")+facet_wrap(~Species)+theme_bw()+theme(axis.title=element_text(color="Blue",face="bold"),plot.title=element_text(color="Green",face="bold"),plot.subtitle=element_text(color="Pink"),panel.grid=element_blank() )```Themes can be saved and reused.```{r}#| fig-height: 4#| fig-width: 6newtheme <-theme(axis.title=element_text(color="Blue",face="bold"),plot.title=element_text(color="Green",face="bold"),plot.subtitle=element_text(color="Pink"),panel.grid=element_blank())ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Sepal.Width))+geom_smooth(method="lm")+scale_color_continuous(name="New Legend Title")+scale_x_continuous(breaks=1:8)+labs(title="This Is A Title",subtitle="This is a subtitle",x=" Petal Length", y="Petal Width", caption="This is a little caption.")+facet_wrap(~Species)+theme_bw()+ newtheme```### Controlling legendsHere we see two legends based on the two aesthetic mappings.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species,size=Sepal.Width))```If we don't want to have the extra legend, we can turn off legends individually by aesthetic.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species,size=Sepal.Width))+guides(size="none")```We can also turn off legends by geom.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species,size=Sepal.Width),show.legend=FALSE)```Legends can be moved around using theme.```{r}#| fig-height: 4#| fig-width: 6ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species,size=Sepal.Width))+theme(legend.position="top",legend.justification="right")```Legend rows can be controlled in a finer manner.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species,size=Sepal.Width))+guides(size=guide_legend(nrow=2,byrow=TRUE),color=guide_legend(nrow=3,byrow=T))+theme(legend.position="top",legend.justification="right")```### LabellingItems on the plot can be labelled using the `geom_text` or `geom_label` geoms.```{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species))+geom_text(aes(label=Species,hjust=0),nudge_x=0.5,size=3)``````{r}#| fig-height: 4#| fig-width: 5ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species))+geom_label(aes(label=Species,hjust=0),nudge_x=0.5,size=3)```The R package `ggrepel` allows for non-overlapping labels.```{r}#| fig-height: 4#| fig-width: 6library(ggrepel)ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species))+geom_text_repel(aes(label=Species),size=3)```### AnnotationsCustom annotations of any geom can be added arbitrarily anywhere on the plot.```{r}#| fig-height: 4#| fig-width: 6ggplot(data=iris,mapping=aes(x=Petal.Length,y=Petal.Width))+geom_point(aes(color=Species))+annotate("text",x=2.5,y=2.1,label="There is a random line here")+annotate("segment",x=2,xend=4,y=1.5,yend=2)```### Barplots```{r}#| fig-height: 4#| fig-width: 6ggplot(data=iris,mapping=aes(x=Species,y=Petal.Width))+geom_bar(stat="identity")```### Flip axesx and y axes can be flipped using `coord_flip`.```{r}#| fig-height: 4#| fig-width: 6ggplot(data=iris,mapping=aes(x=Species,y=Petal.Width))+geom_bar(stat="identity")+coord_flip()```### Error BarsAn example of using error bars with points. The mean and standard deviation is computed. This is used to create upper and lower bounds for the error bars.```{r}#| fig-height: 4#| fig-width: 6dfr <- iris %>%group_by(Species) %>%summarise(mean=mean(Sepal.Length),sd=sd(Sepal.Length)) %>%mutate(high=mean+sd,low=mean-sd)ggplot(data=dfr,mapping=aes(x=Species,y=mean,color=Species))+geom_point(size=4)+geom_errorbar(aes(ymax=high,ymin=low),width=0.2)```## 2. Economist ScatterplotThe aim of this challenge is to recreate the plot below originally published in [The Economist](https://www.economist.com/blogs/dailychart/2011/12/corruption-and-development). The graph is a scatterplot showing the relationship between *Corruption Index* and *Human Development Index* for various countries.### DataDownload the data csv file.<a class="btn btn-secondary btn-sm" href="https://www.dropbox.com/s/7imn0eoey9ckxh5/data_economist.csv?dl=1" role="button">{{< fa download >}} data_economist.csv</a>Start by reading in the data.```{r}ec <-read.csv("assets/data_economist.csv",header=T)head(ec)```Make sure that the fields are of the correct type. The x-axis field 'CPI' and the y-axis field 'HDI' must be of numeric type. The categorical field 'Region' must be of Factor type.```{r}str(ec)```We need to first modify the region column. The current levels in the 'Region' field are:```{r}levels(ec$Region)```But, the categories on the plot are different and need to be changed as follows:``` From ToEU W. Europe OECDAmericas AmericasAsia Pacific Asia & OceaniaEast EU Cemt Asia Central & Eastern EuropeMENA Middle East & North AfricaSSA Sub-Saharan Africa```Since the 'To' strings are a bit too long to be in one line on the legend, use `\n` to break a line into two lines.::: {.callout-tip}`\n` is the newline character in R.:::``` From ToEU W. Europe OECDAmericas AmericasAsia Pacific Asia &\nOceaniaEast EU Cemt Asia Central &\nEastern EuropeMENA Middle East &\nNorth AfricaSSA Sub-Saharan\nAfrica```The strings can be renamed using string replacement or substitution. But a easier way to do it is to use `factor()`. The arguments `levels` and `labels` in function `factor()` can be used to rename factors.```{r}ec$Region <-factor(ec$Region,levels =c("EU W. Europe","Americas","Asia Pacific","East EU Cemt Asia","MENA","SSA"),labels =c("OECD","Americas","Asia &\nOceania","Central &\nEastern Europe","Middle East &\nNorth Africa","Sub-Saharan\nAfrica"))```Our new Regions should look like:```{r}levels(ec$Region)```### PointsStart building up the basic plot.::: {.callout-tip}Provide data.frame 'ec' as the data and map field 'CPI' to the x-axis and 'HDI' to the y-axis. Use `geom_point()` to draw point geometry. To select shapes, see [here](https://www.google.se/search?q=r+pch&oq=R+pch). Circular shape can be drawn using 1, 16, 19, 20 and 21. Using shape '21' allows us to control stroke color, fill color and stroke thickness for the points. Check out `?geom_point` and look under 'Aesthetics' for the various possible aesthetic options. Set shape to 21, size to 3, stroke to 0.8 and fill to white.:::```{r}#| fig-height: 4#| fig-width: 7ggplot(ec,aes(x=CPI,y=HDI,color=Region))+geom_point(shape=21,size=3,stroke=0.8,fill="white")```Notice how '\n' has created newlines in the Legend.### TrendlineNow, we add the trend line using `geom_smooth`. Check out `?geom_smooth` and look under 'Arguments' for argument options and 'Aesthetics' for the aesthetic options.- Use method 'lm' and use a custom formula of `y~poly(x,2)` to approximate the curve seen on the plot. Turn off confidence interval shading. Set line thickness to 0.6 and line color to red.```{r}#| fig-height: 4#| fig-width: 7ggplot(ec,aes(x=CPI,y=HDI,color=Region))+geom_point(shape=21,size=3,stroke=0.8,fill="white")+geom_smooth(method="lm",formula=y~poly(x,2),se=F,size=0.6,color="red")```Notice that the line in drawn over the points due to the plotting order. We want the points to be over the line. So reorder the geoms. Since we provided no aesthetic mappings to `geom_smooth`, there is no legend entry for the trendline. We can fake a legend entry by providing an aesthetic, for example; `aes(fill="red")`. We do not use the color aesthetic because it is already in use and would give us reduced control later on to modify this legend entry.```{r}#| fig-height: 4#| fig-width: 7p <-ggplot(ec,aes(x=CPI,y=HDI,color=Region))+geom_smooth(aes(fill="red"),method="lm",formula=y~poly(x,2),se=F,color="red",size=0.6)+geom_point(shape=21,size=3,stroke=0.8,fill="white")p```### Text LabelsNow we add the text labels. Only a subset of countries are plotted. The list of countries to label is shown below.``` "Congo","Afghanistan","Sudan","Myanmar","Iraq","Venezuela","Russia","Argentina","Brazil","Italy","South Africa","Cape Verde","Bhutan","Botswana","Britian","New Zealand","Greece","China","India","Rwanda","Spain","France","United States","Japan","Norway","Singapore","Barbados","Germany"```- Use `geom_text` to subset the original data.frame to the reduced set above and plot the labels as text. See `?geom_text`.```{r}#| fig-height: 5#| fig-width: 7labels <-c("Congo","Afghanistan","Sudan","Myanmar","Iraq","Venezuela","Russia","Argentina","Brazil","Italy","South Africa","Cape Verde","Bhutan","Botswana","Britian","New Zealand","Greece","China","India","Rwanda","Spain","France","United States","Japan","Norway","Singapore","Barbados","Germany")p+geom_text(data=subset(ec,Country %in% labels),aes(label=Country),color="black")```### Custom FontCustom font can be used for the labels by providing the font name to argument `family` like so `geom_text(family="fontname")`. If you do not want to bother with fonts, just avoid the `family` argument in `geom_text` and skip this part.Using custom fonts can be tricky business. To use a font name, it must be installed on your system and it should be imported into the R environment. This can be done using the `extrafont` package. Try importing one of the fonts available on your system. Not all fonts work. `extrafont` prefers **.ttf** fonts. If a font doesn't work, try another.```{r}#| eval: falselibrary(extrafont)font_import(pattern="Gidole",prompt=FALSE)# load fonts for pdfloadfonts()# list available fonts in Rfonts()```The actual font used on the Economist graph is something close to [ITC Officina Sans](https://www.fontshop.com/families/itc-itc-officina-sans). Since this is not a free font, I am using a free font called [Gidole](https://gidole.github.io/).```{r}#| fig-height: 5#| fig-width: 7p+geom_text(data=subset(ec,Country %in% labels),aes(label=Country),color="black",family="Gidole")```### Label OverlapTo avoid overlapping of labels, we can use a `ggplot2` extension package `ggrepel`. We can use function `geom_text_repel()` from the `ggrepel` package. `geom_text_repel()` has the same arguments/aesthetics as `geom_text` and a few more. Skip the `family=Gidole` part if you do not want to change the font.```{r}#| fig-height: 5#| fig-width: 7library(ggrepel)p <- p+geom_text_repel(data=subset(ec,Country %in% labels),aes(label=Country),color="black",box.padding=unit(1,'lines'),segment.size=0.25,size=3,family="Gidole")p```### AxesNext step is to adjust the axes breaks, axes labels, point colors and relabeling the trendline legend text.- Change axes labels to 'Corruption Perceptions Index, 2011 (10=least corrupt)' on the x-axis and 'Human Development Index, 2011 (1=best)' on the y-axis. Set breaks on the x-axis from 1 to 10 by 1 increment and y-axis from 0.2 to 1.0 by 0.1 increments.```{r}#| fig-height: 5#| fig-width: 7p <- p+scale_x_continuous(name="Corruption Perceptions Index, 2011 (10=least corrupt)",breaks=1:10,limits=c(1,10))+scale_y_continuous(name="Human Development Index, 2011 (1=best)",breaks=seq(from=0,to=1,by=0.1),limits=c(0.2,1))p```### Scale ColorsNow we want to change the color palette for the points and modify the legend text for the trendline.- Use `scale_color_manual()` to provide custom colors. These are the colors to use for the points: `"#23576E","#099FDB","#29B00E", "#208F84","#F55840","#924F3E"`. - Use `scale_fill_manual` to change the trendline label since it's a fill scale. The legend entry for the trendline should read 'R^2=52%'.```{r}#| fig-height: 5#| fig-width: 7p <- p+scale_color_manual(values=c("#23576E","#099FDB","#29B00E", "#208F84","#F55840","#924F3E"))+scale_fill_manual(name="trend",values="red",labels=expression(paste(R^2,"=52%")))p```### TitleTitle and caption can be added with `labs`.- Set the title to 'Corruption and human development'. - Set the caption to 'Sources: Transparency International; UN Human Development Report'.```{r warning=FALSE}#| fig-height: 5#| fig-width: 7p <- p+labs(title="Corruption and human development", caption="Sources: Transparency International; UN Human Development Report")p```### ThemeWe want to move the legend to the top and as a single row. This can be done using `theme()` option `legend.position`. See `?theme`. `guides()` is used to set the number of rows to 1. We also set a custom font for all text elements using `base_family="Gidole"`. This can be skipped if a font change is not required.```{r warning=FALSE}#| fig-height: 5#| fig-width: 7p <- p+guides(color=guide_legend(nrow=1))+ theme_bw(base_family="Gidole")+ theme(legend.position="top")p```Now we do some careful refining with themes.- Turn off minor gridlines- Turn off major gridlines on x-axis- Remove the gray background- Remove panel border- Remove legend titles- Make axes titles italic- Turn off y-axis ticks- Change x-axis ticks to color grey60- Make plot title bold- Decrease size of caption to size 8```{r warning=FALSE}#| fig-height: 5#| fig-width: 7p+theme(panel.grid.minor=element_blank(), panel.grid.major.x=element_blank(), panel.background=element_blank(), panel.border=element_blank(), legend.title=element_blank(), axis.title=element_text(face="italic"), axis.ticks.y=element_blank(), axis.ticks.x=element_line(color="grey60"), plot.title=element_text(face="bold"), plot.caption=element_text(hjust=0,size=8))```And now our plot is ready and we can compare with the original. Close? Close enough?The full script for this challenge is summarized here:```{r}#| eval: false# read dataec <-read.csv("assets/data_economist.csv",header=T)# refactorec$Region <-factor(ec$Region,levels =c("EU W. Europe","Americas","Asia Pacific","East EU Cemt Asia","MENA","SSA"),labels =c("OECD","Americas","Asia &\nOceania","Central &\nEastern Europe","Middle East &\nNorth Africa","Sub-Saharan\nAfrica"))# labelslabels <-c("Congo","Afghanistan","Sudan","Myanmar","Iraq","Venezuela","Russia","Argentina","Brazil","Italy","South Africa","Cape Verde","Bhutan","Botswana","Britian","New Zealand","Greece","China","India","Rwanda","Spain","France","United States","Japan","Norway","Singapore","Barbados","Germany")# plottingp1 <-ggplot(ec,aes(x=CPI,y=HDI,color=Region))+geom_smooth(aes(fill="red"),method="lm",formula=y~poly(x,2),se=F,color="red",size=0.6)+geom_point(shape=21,size=3,stroke=0.8,fill="white")+geom_text_repel(data=subset(ec,Country %in% labels),aes(label=Country),color="black",box.padding=unit(1,'lines'),segment.size=0.25,size=3,family="Gidole")+scale_x_continuous(name="Corruption Perceptions Index, 2011 (10=least corrupt)",breaks=1:10,limits=c(1,10))+scale_y_continuous(name="Human Development Index, 2011 (1=best)",breaks=seq(from=0,to=1,by=0.1),limits=c(0.2,1))+scale_color_manual(values=c("#23576E","#099FDB","#29B00E", "#208F84","#F55840","#924F3E"))+scale_fill_manual(name="trend",values="red",labels=expression(paste(R^2,"=52%")))+labs(title="Corruption and human development",caption="Sources: Transparency International; UN Human Development Report")+guides(color=guide_legend(nrow=1))+theme_bw(base_family="Gidole")+theme(legend.position="top",panel.grid.minor=element_blank(),panel.grid.major.x=element_blank(),panel.background=element_blank(),panel.border=element_blank(),legend.title=element_blank(),axis.title=element_text(face="italic"),axis.ticks.y=element_blank(),axis.ticks.x=element_line(color="grey60"),plot.title=element_text(face="bold"),plot.caption=element_text(hjust=0,size=8))p1```## 3. WSJ HeatmapThe aim of this challenge is to recreate the plot below originally published in [The Wall Street Journal](http://graphics.wsj.com/infectious-diseases-and-vaccines/). The plot is a heatmap showing the normalized number of cases of measles across 51 US states from 1928 to 2003. X-axis shows years and y-axis shows the names of states. The color of the tiles denote the number of measles cases per 100,000 people. Introduction of the measles vaccine is shown as the black line in 1963.### DataDownload the data csv file.<a class="btn btn-secondary btn-sm" href="https://www.dropbox.com/s/19p8vku0i9np26b/data_wsj.csv?dl=1" role="button">{} data_wsj.csv</a>Start by reading in the data. This .csv file has two lines of comments so we need to skip 2 lines while reading in the data. We also add `stringsAsFactors=F` to avoid the automatic conversion of character fields to factor type.```{r}me <-read.csv("assets/data_wsj.csv",header=T,stringsAsFactors=F,skip=2)head(me)```Check the data type for the fields.```{r}str(me)```Looking at this dataset, there is going to be quite a bit of data clean-up and tidying before we can plot it. Here are the steps we need to take:- The data needs to be transformed to long format.- Replace all "-" with NAs- The number of cases across each state is a character and needs to be converted to numeric- Collapse (sum) week-level data to year.- Abbreviate state names### Tidy DataConvert the wide format to long format using the function `gather()` from package `dplyr`.```{r}me1 <- me %>%gather(key=state,value=value,-YEAR,-WEEK)head(me1)```Now, replace all '-' with NA in the field value. We use the function `str_replace()` from R package `stringr`. Then convert the value field to numeric.```{r}me2 <- me1 %>%mutate(value=str_replace(value,"^-$",NA_character_),value=as.numeric(value))head(me2)```Sum up the week-level information to year-level information. This means rather than having``` YEAR WEEK state value1 1928 1 ALABAMA 3.672 1928 2 ALABAMA 6.253 1928 3 ALABAMA 7.95...5501 1957 41 ALASKA 2.165502 1957 42 ALASKA 0.435503 1957 43 ALASKA 1.30...```we should have one value per year per state.``` YEAR state value1 1928 ALABAMA 3.672 1929 ALABAMA 3.20...5501 1957 ALASKA 2.165502 1958 ALASKA 2.05...```The solution is to sum up all the cases for a state for all weeks within a year into one value for that year. This can be done using the `summarise()` function from package `dplyr`.- A custom function is used to sum over weeks. If all values are NA, then result is NA. If some values are NA, the NAs are removed and the remaining numbers are summed.- The dots in state names are replaced by spaces and the words are converted to title case (First letter capital and rest lowercase). - We also convert the column names to lowercase for consistency.```{r}fun1 <-function(x) ifelse(all(is.na(x)),NA,sum(x,na.rm=TRUE))me3 <- me2 %>%group_by(YEAR,state) %>%summarise(total=fun1(value)) %>%mutate(state=str_replace_all(state,"[.]"," "),state=str_to_title(state))colnames(me3) <-tolower(colnames(me3))head(me3)``````{r}str(me3)```The data is now ready for plotting.### TileWe can build up a basic ggplot and heatmap tiles can be plotted using the geom `geom_tile`. 'year' is mapped to the x-axis, 'state' to the y-axis and fill color for the tiles is the 'total' value.```{r}#| fig-height: 6#| fig-width: 9ggplot(me3,aes(x=year,y=state,fill=total))+geom_tile()```Add borders around the tiles. We use `reorder(state,desc(state))` to reverse the order of states so that it reads A-Z from top to bottom.```{r}#| fig-height: 6#| fig-width: 9p <-ggplot(me3,aes(x=year,y=reorder(state,desc(state)),fill=total))+geom_tile(color="white",size=0.25)p```### ScalesThe extra space on left and right (gray) of the plot is removed using argument `expand` in `scales`. X-axis breaks are redefined at 10 year intervals from 1930 to 2010. Custom colors are used for the tiles: `"#e7f0fa","#c9e2f6","#95cbee","#0099dc","#4ab04a", "#ffd73e","#eec73a","#e29421","#f05336","#ce472e"`. Since the color scale is a fill color on a continuous value and we want to supply n new colors, we use `scale_fill_gradientn`. Tiles with missing value is set to the color `"grey90"`.```{r}#| fig-height: 6#| fig-width: 9cols <-c("#e7f0fa","#c9e2f6","#95cbee","#0099dc","#4ab04a", "#ffd73e","#eec73a","#e29421","#f05336","#ce472e")p +scale_y_discrete(expand=c(0,0))+scale_x_continuous(expand=c(0,0),breaks=seq(1930,2010,by=10))+scale_fill_gradientn(colors=cols,na.value="grey95")```The fill scale can be further refined to resemble that of the original plot.```{r}#| fig-height: 6#| fig-width: 9cols <-c("#e7f0fa","#c9e2f6","#95cbee","#0099dc","#4ab04a", "#ffd73e","#eec73a","#e29421","#f05336","#ce472e")p <- p+scale_y_discrete(expand=c(0,0))+scale_x_continuous(expand=c(0,0),breaks=seq(1930,2010,by=10))+scale_fill_gradientn(colors=cols,na.value="grey95",limits=c(0,4000),values=c(0,0.01,0.02,0.03,0.09,0.1,0.15,0.25,0.4,0.5,1),labels=c("0k","1k","2k","3k","4k"),guide=guide_colourbar(ticks=T,nbin=50,barheight=.5,label=T, barwidth=10))p```### TitleWe can remove the x and y axes titles and add a plot title.```{r}#| fig-height: 6#| fig-width: 9p <- p+labs(x="",y="",fill="",title="Measles")p```### Fixed CoordsWe can use `coord_fixed()` to fix the coordinates for equal values in x and y direction. This should render perfectly square tiles.```{r}#| fig-height: 6#| fig-width: 9p <- p+coord_fixed()p```### AnnotationAdd the annotation line and text to denote the introduction of the vaccine. The line is at the position 1963. Custom font 'Gidole' is used here. This can be skipped.```{r}#| fig-height: 6#| fig-width: 9p <- p+geom_segment(x=1963,xend=1963,y=0,yend=51.5,size=.6,alpha=0.7) +annotate("text",label="Vaccine introduced",x=1963,y=53, vjust=1,hjust=0,size=I(3),family="Gidole")p```### ThemeHere we change the following aspects of the plot using `theme`:- Change theme to `theme_minimal` to remove unnecessary plot elements.- Optional custom font. See 'Custom font' section under 'Economist Scatterplot'. - Position the legend to bottom center.- Set legend font to color grey20.- Adjust size and justification of x and y axes text- Adjust title justification- Remove all gridlines```{r warning=FALSE}#| fig-height: 6#| fig-width: 9p+theme_minimal(base_family="Gidole")+ theme(legend.position="bottom", legend.justification="center", legend.direction="horizontal", legend.text=element_text(color="grey20"), axis.text.y=element_text(size=6,hjust=1,vjust=0.5), axis.text.x=element_text(size=8), axis.ticks.y=element_blank(), title=element_text(hjust=-.07,vjust=1), panel.grid=element_blank())```Our plot is ready and we can compare it to the original version.The full code for this challenge is here:```{r}#| eval: false# custom summing functionfun1 <-function(x) ifelse(all(is.na(x)),NA,sum(x,na.rm=TRUE))# read datame3 <-read.csv("assets/data_wsj.csv",header=T,stringsAsFactors=F,skip=2) %>%gather(key=state,value=value,-YEAR,-WEEK) %>%mutate(value=str_replace(value,"^-$",NA_character_),value=as.numeric(value)) %>%group_by(YEAR,state) %>%summarise(total=fun1(value)) %>%mutate(state=str_replace_all(state,"[.]"," "),state=str_to_title(state))colnames(me3) <-tolower(colnames(me3))# custom colorscols <-c("#e7f0fa","#c9e2f6","#95cbee","#0099dc","#4ab04a", "#ffd73e","#eec73a","#e29421","#f05336","#ce472e")# plottingp <-ggplot(me3,aes(x=year,y=reorder(state,desc(state)),fill=total))+geom_tile(color="white",size=0.25)+scale_y_discrete(expand=c(0,0))+scale_x_continuous(expand=c(0,0),breaks=seq(1930,2010,by=10))+scale_fill_gradientn(colors=cols,na.value="grey95",limits=c(0,4000),values=c(0,0.01,0.02,0.03,0.09,0.1,0.15,0.25,0.4,0.5,1),labels=c("0k","1k","2k","3k","4k"),guide=guide_colourbar(ticks=T,nbin=50,barheight=.5,label=T, barwidth=10))+labs(x="",y="",fill="",title="Measles")+coord_fixed()+geom_segment(x=1963,xend=1963,y=0,yend=51.5,size=.9) +annotate("text",label="Vaccine introduced",x=1963,y=53, vjust=1,hjust=0,size=I(3),family="Gidole")+theme_minimal(base_family="Gidole")+theme(legend.position=c(.5,-.13),legend.direction="horizontal",legend.text=element_text(color="grey20"),plot.margin=grid::unit(c(.5,0,1.5,0),"cm"),axis.text.y=element_text(size=6,hjust=1,vjust=0.5),axis.text.x=element_text(size=8),axis.ticks.y=element_blank(),panel.grid=element_blank(),title=element_text(hjust=-.07,vjust=1),panel.grid=element_blank())```#### Acknowledgements:• This document was adapted from [RaukR-2023](https://nbisweden.github.io/raukr-2023/labs/ggplot/),• shared under [GPL-3 License](https://choosealicense.com/licenses/gpl-3.0/)• [SLUBI](https://www.slubi.se/)